Finding Causative Mutations With A Candidate SNP Approach
Why Do We Need Mutational Genomics?
A fork is not a hairbrush - it just has some similar properties
Genomics has come a long way. We can now sequence genomes quickly and to a reasonable degree of accuracy such that with sequence based approaches we can create in a high-throughput manner an inventory of sub-regions in a genome that we think are genes. We know the functions (or some of the functions) of lots of genes and we can infer functions of newly discovered genes by comparison of sequence or structure, basically by seeing whether our new thing looks like something else.
But these methods are actually only PREDICTIONS of function. Looking a bit like something else is only a clue to what something does. It frequently fails us.

Ariel thinks the fork is a brush, it does look like one…
So finding the function of a gene isn’t straightforward
What makes things worse is that often we don’t start with the gene itself. We start with some biological process and want to find genes with a role in that process. Because our functional predictions aren’t perfect we can’t collect all the genes and start trying them out one-by-one.

One of these is the key to the bathroom. I hope that you’re not desperate!
We need something smarter - mutational genomics is this smarter thing
With mutational genomics we deal initially with the effect of the gene on the whole organism. By performing mutagenesis on our favourite organism then carrying out a screen that selects individuals that have changed in the phenotype we are interested in, we have our first foothold. We can study those individuals and apply the principles of genetics, use modern high-throughput sequencing and bioinformatics tools to identify the gene causing that phenotype change (or at least ones involved in the process we have messed up).
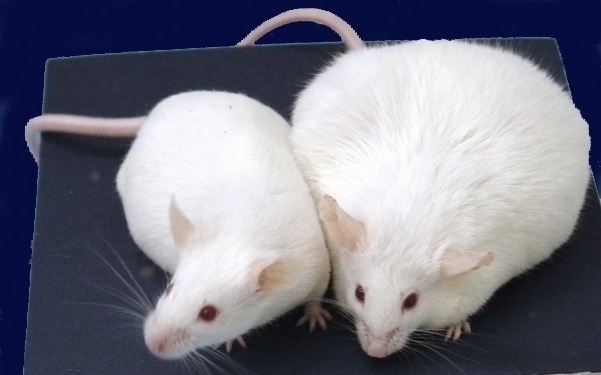
One of these mice is not like the other mice - it has a phenotype change
Our objective
This will be the focus of this workshop - how to go from samples identified in a genetic screen to a short-list of candidate genes using Galaxy tools and software.

